|
Looking for help with Power Pivot in Excel 2013? Go to on Office.com. After you have created a connection to an external data source, you can later modify that connection in these ways: • You can change the connection information, including the file, feed, or database used as a source, its properties, or other provider-specific connection options. • You can change table and column mappings, and remove references to columns that are no longer used. • You can change the tables, views, or columns that you get from the external data source. For more information, see.  
• In the PowerPivot window, click the Home tab, and in the Connections group, click Existing Connections. • Select the current database connection and click Edit. For this example, the Table Import Wizard opens to the page that configures an Access database. However, depending on the type of data source you are changing, the provider might be different, and also the properties that are available. • In the Edit Connection dialog box, click Browse to locate another database of the same type but with a different name or location. As soon as you change the database file, a message appears indicating that you need to save and refresh the tables in order to see the new data. • Click Save, and then click Close. • On the Home tab, in the Get External Data group, click Refresh, and then click Refresh All. 
The tables are refreshed using the new data source, but with the original data selections. • In the PowerPivot window, click the Design tab, and in the Properties group, click Table Properties. The name of the current workbook table is displayed in the Table Name box. The Source Name box contains the name of the table in the external data source. If columns are named differently in the source and in the workbook, you can toggle between the two sets of column names by selecting the options Source or PowerPivot data (workbook). • To change the table that is used as a data source, for Source Name, select a different table than the current one. • Change column mappings if needed: • To add columns that are present in the source but not in the workbook, select the checkbox beside the column name. The actual data will be loaded into the workbook the next time you refresh. • If some columns in the workbook are no longer available in the current data source, a message appears in the notification area that lists the invalid columns. You do not need to do anything else. Excel Edit Properties Software antivirus report. This download is virus-free. This file was last analysed by Free Download Manager Lib 10 days ago. Excel Document Details Editor is a tool devised to culminate the workload of altering details of Excel files. The software has the latest characteristics rigged in it. MS Excel File Properties Changer, as its name suggests, is a prograù that allows editing the properties of an Excel spreadsheet such as summary information, file. Register for Exam 77-420 and view official preparation materials to get hands-on experience with Excel 2013. Sep 09, 2016 We have no issues editing document properties in a new or existing Excel document as long as it is NOT shared. If you share the document, which happens. • Click Save to apply the changes to your workbook. When you save the current set of table properties, any invalid columns are automatically removed and new columns are added. A message appears indicating that you need to refresh the tables. Click Refresh to load updated data into your workbook. 
0 Comments
====================================================================== Download Sothink iPad iPod iPhone Suite last version for windows 7, 8.1 x32 bit from the or ====================================================================== Form.AutoSize Property (System.Windows. Forms) - MSDN - Microsoft4 Jul 2013. Joy RingTone Converter is a program that lets you create excellent ringtones with the audio formats used by most mobile phones available. Flip PDF Professional is a useful utility to create Flash-based eBooks with a real, life-like page-flipping effect. With it, you are able to edit imported PDF pages by.vb. Net - How do I prevent a form from being resized by the user. Softgroup.Net Forms Resize Softgroup.Net Forms Resize.Net Forms Resize.Net Forms Resize Softgroup.Net Forms Resize is a fast, small and lightweight.NET component that gives your Visual Studio applications resolution independence. Softgroup.Net Forms Resize.Net Forms Resize Softgroup.Net Forms Resize is a fast, small and lightweight.NET component that gives your Visual Studio applications resolution independence. Softgroup.Net Forms Resize supports standard.Net Windows Forms, MDI child and MDI parent Forms and includes DPIAutoResize feature to make your application DPI resolution independent too. This resizer component can be easily added to already designed forms with 1 line of code. Softgroup.Net Forms Resize automatically resize all controls and fonts contained in a.Net Windows Form as they are sized. Softgroup.Net Forms Resize is a fast, small and lightweight.NET component that gives your applications resolution independence. Softgroup.Net Forms Resize supports standard.Net Windows Forms, MDI child and MDI parent Forms and includes DPIAutoResize feature to make your application DPI resolution independent too. Softgroup.Net Forms Resize automatically resize all controls and fonts contained in a.Net Windows Form as they are sized. Only one line of code is needed in the Form_Load event. Screenshot Gallery Softgroup.Net Forms Resize. Why do we need Blu-ray Creator? Blu-ray Creator gives title authors the ability to combine high definition video and audio with interactive menus and internet connectivity, proof their title as they work, and output fully formatted Blu-ray pre-masters for replication. Blu-ray creator software is a kind of software to create blu-ray, by which you can convert some video fotmats such as DVD,AVI, DivX, XviD, VCD, WMV, MPEG4, TD,TS to blu-ray format.Then it can be saved on a hard disk or be burned on a blu-ray disk. Hope this review will help you to choose.  
WinX HD Video Converter Deluxe نرم افزاری تبدیل فرمت فیلم HD حرفه ای برای تبدیل انواع فایل های ویدئویی است. Download the free trial version below to get started. Double-click the downloaded file to install the software. Attain the best Sothink DVD EZWorkshop + iPod Video Suite promo code deals coming from the leader of Software coupons. IPad iPod iPhone Suite Promo Codes.  
If you've got a soft spot for the disco heyday, we'd guess this screensaver will suit your tastes. In 3D Disco Girl, a young woman gyrates across a lit-up dance floor, as a mirror behind her shows three reflections of her figure. We like the colorful graphics, although the movement of the girl isn't completely realistic. 3D Disco Girl Download 3D Disco Girl dresses in slacks and bare feet and invokes a feel of the '70s and '80s. The disco scene has a flashing multicolored floor. 
If you like the program's appearance but can't stand the disco soundtrack, you always can add your own music. You also can tweak the screensaver's brightness and RGB color levels. On the downside, the demo period is only one short week, and the trial version also displays a nag screen. Nevertheless, we can see how 3D Disco Girl would make a good screensaver for parties or any other places where folks are boogying down. From 3D Disco Girl dresses in slacks and bare feet and invokes a feel of the '70s and '80s. The disco scene has a flashing multicolored floor, changing background, and disco girls dancing in bubbles in the background. The demo contains a full set of features. Have fun customizing this screensaver to your preferences. In addition to background tint, separate animation tint, brightness, sound mute, and volume control, you also can play your own MP3 music. The application contains animated GIF images and sound files, and password protection is available. 
3D Disco Girl is a Desktop Utilities software developed by Unique 3D Screensavers. After our trial and test, the software is proved to be official, secure and free. Here is the official description for 3D Disco Girl: Edit By BS Editor: 3D Girl in slacks and bare feet invokes a feel of the 70s and 80s. Free porn movies from the most popular XXX tubes. Watch daily updated stream porn movies online! Only at HHJCC hot Cartoon, 3d Sister, 3d Slave, 3d Space, Hot. Dance, disco, party the night away! Style this fun party goer in colorful, crop top and cut off everything! Throw in some cute boots and candy accessories for a. The disco scene has a flashing multi-colored floor, changing background and disco girls dancing in bubbles in the background. Demo contains full set of features. Have fun customizing this screen saver to your preferences. In addition to background, separate animation tint, brightness, sound mute and you can play your own mp3 music. 
Int J Infect Dis. 2015 Mar;32:94-100. Doi: 10.1016/j.ijid.2015.01.014. The complex evolution of antibiotic resistance in Mycobacterium tuberculosis. Fonseca JD(1), Knight GM(2), McHugh TD(3). Author information: (1)Centre for Clinical Microbiology, University College London, London, NW3 2PF, UK. Electronic address:. Feb 23, 2010. First principles calculations, employed to address the properties of polycrystalline graphene, indicate that the electronic structure of tilt grain boundaries in this system displays a rather complex evolution toward graphene bulk, as the tilt angle decreases, with the generation of a Dirac point, at the Fermi level,. Abstract Recent phylogenomic studies have failed to conclusively resolve certain branches of the placental mammalian tree, despite the evolutionary analysis of genomic data from 32 species. Previous analyses of single genes and retroposon insertion data yielded support for different phylogenetic scenarios for the most basal divergences. The results indicated that some mammalian divergences were best interpreted not as a single bifurcating tree, but as an evolutionary network. In these studies the relationships among some orders of the super-clade Laurasiatheria were poorly supported, albeit not studied in detail. Therefore, 4775 protein-coding genes (6,196,263 nucleotides) were collected and aligned in order to analyze the evolution of this clade. 
Additionally, over 200,000 introns were screened in silico, resulting in 32 phylogenetically informative long interspersed nuclear elements (LINE) insertion events. The present study shows that the genome evolution of Laurasiatheria may best be understood as an evolutionary network. Thus, contrary to the common expectation to resolve major evolutionary events as a bifurcating tree, genome analyses unveil complex speciation processes even in deep mammalian divergences. We exemplify this on a subset of 1159 suitable genes that have individual histories, most likely due to incomplete lineage sorting or introgression, processes that can make the genealogy of mammalian genomes complex. These unexpected results have major implications for the understanding of evolution in general, because the evolution of even some higher level taxa such as mammalian orders may sometimes not be interpreted as a simple bifurcating pattern. Citation: Hallström BM, Schneider A, Zoller S, Janke A (2011) A Genomic Approach to Examine the Complex Evolution of Laurasiatherian Mammals. PLoS ONE 6(12): e28199. Editor: Ed Louis, University of Nottingham, United Kingdom Received: August 4, 2011; Accepted: November 3, 2011; Published: December 2, 2011 Copyright: © 2011 Hallström et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. Funding: The present study was funded by the research funding programme “ LOEWE - Landes -Offensive zur Entwicklung Wissenschaftlich-ökonomischer Exzellenz” of Hesse's Ministry of Higher Education, Research, and the Arts. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. Competing interests: The authors have declared that no competing interests exist. Introduction While the placental mammalian tree is becoming increasingly better resolved, it has proven difficult to fully resolve several branches of it as a bifurcating tree, despite the availability and analyses of whole genome data,. While the sheer amount of genomic data should be sufficient to resolve very short branches within the placental mammalian tree, the support for some branches is often ambigious. Interestingly, these problematic branches are characterized by rapid divergences within 1–3 million years (Myr). This makes it possible that speciation related processes, such as incomplete lineage sorting or introgression, lead to gene trees that differ from the species tree,. The complex pattern of retroposon insertion data for the earliest placental mammalian divergences corroborate this idea, suggesting that a network-like evolution instead of a bifurcating tree best depict and interpret the evolutionary process. Other such problematic relationships among placental mammals have been identified by phylogenomic, and retroposon insertion data. 
A case in point is the evolution of the mammalian clade Laurasiatheria, which comprises several orders of placental mammals. Laurasiatheria include the classical orders Perissodactyla, Carnivora, Pholidota, Artiodactyla, Cetacea, Chiroptera, and Lipotyphla,. Initially, morphological and early molecular studies spread these orders to different parts of the mammalian tree or left their position unresolved. More detailed molecular phylogenetic studies grouped these diverse orders into one clade, Laurasiatheria. Early mitogenomic studies suggested a close relationship between carnivores and perissodactyls and this group in turn joined Cetartiodactyla,. Later mitogenomic studies added Chiroptera and parts of a then paraphyletic Lipotyphla to the Laurasiatheria clade, whereas analysis of nuclear genes placed all of Lipotyphla within Laurasiatheria. Finally, the Pholidota (pangolin) were joined with the carnivores by nuclear and mitogenomic studies,. Currently molecular phylogenetic studies generally agree on a (Chiroptera,(Cetartiodactyla, (Perissodactyla, (Pholidota, Carnivora))) branching order,. With the exception of the Pholidota, which lack large-scale genomic sequence data, recent phylogenomic analyses generally support this topology. However, the relationships remained only poorly supported despite the use of some 3 million nucleotides of sequence data from 3400 protein coding genes. So far only one study using rare genomic events such as data from retroposon insertions has been made to study the relationships within Laurasiatheria. In contrast to sequence-based studies, analyses of three retroposon insertions support the grouping of Chiroptera with the Perissodactyla/Carnivora and a new name for this unexpected clade – Pegasoferae – has been suggested. Yet, this study found one retroposon insertion event that contradicted the Chiroptera plus Perissodactyla/Carnivora grouping. This one retroposon insertion supports the traditional sequence-analyses based placement of Chiroptera. A recent phylogenomic study of mammalian relationships involved all tetrapod species from which whole genome data were available. While it is advantageous to increase taxon sampling, this approach leads to the exclusion of large amounts of sequence data when stringent data collection and alignment strategies are employed,. In addition, the inclusion of distantly related species in the analyses even make it possible that orthologs are misidentified, and thus excluded, as paralogs by overly stringent data retrieval algorithms such as recursive BLAST. In order to specifically analyse laurasiatherian relationships with a dataset maximized for the amount of phylogenetically informative data, only human and mouse are used as outgroups to root the tree in this study. These species have among the best genomic sequence coverage and annotation.  
This paper discusses five of these characteristics and presents a strategy for function optimization called the shuffled complex evolution (SCE) method, which promises to be robust, effective, and efficient for a broad class of problems. The SCE method is based on a synthesis of four concepts that have proved successful for. Furthermore, there is an unequivocal consensus that these two species are joined in the clade Euarchontoglires which is the sister group to Laurasiatheria within Boreotheria. Thus, human and mouse are the ideal outgroups for this study. By also utilizing the recently released genome of the giant panda ( Ailuropoda melanoleuca), this approach allows the collection of a larger number of genes from more species than in previous phylogenomic studies. Therefore, analyses based on concatenated data and single genes allow for a more detailed study of laurasiatherian relationships. In addition, the quality and quantity of the genome data have been steadily improving. This makes in silico searches for phylogenetic informative retroposon insertion data feasible for evaluating hypotheses that were based on sequence data analysis. Long interspersed nuclear elements 1 (LINE 1) retroposon sequences were used for these searches, because these elements were active during this time of placental mammalian evolution and have successfully been used in other phylogenetic studies –. These are currently the only known retroposons that are common to different orders, while short interspersed nuclear elements (SINEs) are order-specific. The names, order and sequence coverage of the species included in this study. Data collection and alignment was, with a few exceptions, performed as described previously and is thus only briefly detailed here. Orthologs were identified with the recursive BLAST method. Sequences were translated to amino acids and aligned using MUSCLE. The resulting alignments were then back-translated to nucleotides. Any alignment showing an overall nucleotide difference larger than 30% between any two species was discarded. As an additional filtering step, uninformative quickly evolving sites were eliminated by the program Noisy, version 1.5.9. Phylogenetic analysis using maximum likelihood was performed using the programs Treefinder (TF) and RAxML 7.0.4, applying the GTR model to nucleotide data and WAG2000 to amino acid data. In both cases, rate heterogeneity was applied using 4 gamma rate categories, 4G+I. Both heuristic searches and exhaustive tree comparisons, under the assumption of monophyletic orders were performed. Divergence times were estimated from overall best amino acid (AA) ML tree using 6 calibration points () and the nonparametric rate smoothing method on a logarithmic scale (NPRS-LOG) as implemented in TF. Codon-based tree reconstruction was performed using the Markov Chain Monte Carlo (MCMC) method implemented in BEAST, using its BEAGLE library for computing on graphics processing units (GPUs). This decreases computation times by a factor of up to 90. The analyses were performed using a semi-parametric codon model based on principal component analysis of mammalian sequence data. For each alignment and topology, the model-parameters as well as the branch lengths were optimized with a chain length of 700,000 sampled every 500 tree. Instead of maximizing the likelihood, BEAST allows an estimation of the marginal log-likelihood (mLogL) by integrating over the whole parameter space,. 
Tree and model comparisons can be performed using the Bayes factor, which can be approximated as the difference of the mLogLs. For 229 trees, BEAST failed to successfully optimize the parameters. These trees were excluded from further analysis. In addition to the analyses of concatenated data, all gene alignments with sequence data from all species were analyzed separately. The problem was reduced to resolving the relationships of four orders, leaving 15 possible topologies that were individually evaluated by ML analyses for each of the 1159 gene alignments. The same models as outlines above were used with parameters estimated from individual alignments. Information from likelihood maximizations on the 15×1159 gene trees were analyzed by counting how often each topology was among the most likely trees and how often a topology was rejected by another topology with a significantly higher likelihood. A significantly higher likelihood is defined as one that is larger than two log-likelihood units from the original. When different topologies had the same likelihood for single-gene alignments, they were counted individually for each tree. In addition, all likelihood values for a given topology and data set were added up in order to compare the total likelihoods of the different topologies. This approach corresponds to the “separate” analysis according to the definition in Pupko et al.,. Since the mLogLs of the codon-based analysis are expected values and not maxima, so typically no two topologies end up with exactly the same mLogL. Thus, a tolerance of 0.5 LogL units was used, and two values that lay within two mLogLs of each other were considered as being equal. A topology was rejected if its mLogL was 10 units lower than the highest. The ML trees from the single-gene analysis were also used to construct a consensus network using the SplitsTree4 program, which is used to illustrate the conflicts of the phylogenetic signal. For the five most likely tree topologies, the influence of several properties of the sequences on the outcome of the codon analysis was tested. The evaluated factors were alignment length, longest distance among the 15 sequences, sum of all pair wise distances, deviation of the codon usage frequencies from the average over all alignments and deviation of the nucleotide usage frequencies from the average. For each factor, the alignments were divided into two equal-sized groups; those with the largest values and those with the smallest values. It was then counted how often each topology was the only one with highest mLogL. Chi-square tests were performed to quantify the significance of the difference between the two “best” distributions of topologies. Finally, a multilocus Bayesian analysis using the program BEST was performed. This method attempts to construct a species tree using a multiple estimated gene trees. This is done by utilizing a Bayesian hierarchical model to combine traditional phylogenetics with coalescent theory. 763 genes (1,313,880 nucleotide characters) were selected for maximum alignment coverage and length. This data set was analyzed in BEST, with all parameters unlinked, runnning for 15,000,000 generations, with two simultaneous runs each with one “heated” and one “cold chain. The first 1.500.000 generations (10%) were discarded as burnin. Retroposon analysys For the retroposon insertion analysis intron sequences longer than 300 bp and shorter than 3000 bp were collected from the Ensembl database (version 49) for the cow, dog, horse and microbat genomes, respectively. Between 40,000 and 95,000 introns were identified in each of the species above. Retroposed elements in these introns were identified using the program RepeatMasker version 3.2.8 (). From all identified repeated elements only LINE1 elements were considered for the search and phylogenetic analysis. In total 47,535 LINE1 elements were identified, of which 22,873 were found in the horse genome, 13,359 in the dog genome, 6,557 in the cow genome and 4,756 in the bat genome. Using these intron sequences the orthologous region in the other three species were identified. The full sequences of orthologous genes were extracted, based on Ensembl orthology data. The relevant intron sequences were located by making local pair wise alignments with 80 bp of exon sequence located upstream and downstream of the intron. In cases where the intron could be located in all four species a four-way multiple sequence alignment was created using MAFFT. This resulted in 19,725 alignments that were guided by 7576 retropson insertions that were initially identified in the horse, 7248 in the dog, 2793 in the cow, and 2108 in the bat, respectively. All four-way alignments were screened for retroposons that were present in either two or three of the species, and absent in the others. These retroposon inserts were considered potential markers for the phylogenetic relationships between the four orders. Finally intron sequences from the remaining laurasiatherian species and, when possible, outgroup sequences were added to the alignments. Additionally, several hundred alignments in which the insertion was present in either only one or all four species were randomly selected and manually screened for potentially informative retroposon markers for other parts of the laurasiatherian tree. The alignments of the, in total 25, informative L1 retroposon insertions that were used for the tree and network are shown in. A consensus tree, was constructed with SplitsTree4 from all partial trees corresponding to the L1 retroposon data for the conflicting hypotheses among the four laurasiatherian orders. The branch lengths are determined by the number of retroposon insertions supporting each topology. Sequence analysis The final alignment consisted of 6,196,263 nucleotide characters (translating to 2,065,421 amino acid characters) from 4775 genes, represented by 12 ingroup species and two outgroup species; human and mouse. Provides a list of all included species and their sequence coverage in the alignment. After eliminating potentially homoplastic sites, the alignment length was reduced to 4,314,195 characters for the nucleotide data and 1,476,398 characters for the amino acid data. The average sequences coverage of the alignment was 85.3%. In heuristic analyses and RAxML parametric bootstrap analysis the relationships within the orders were unanimous, but some inter-ordinal relationships received only limited support. Shows the best-supported tree and branch lengths based on maximum likelihood (ML) analysis of concatenated amino acid (AA) data. This topology was also the best or among the best supported in other analyses. For further evaluating the topology and the support assigned to it, exhaustive analyses were performed on the relationships among the five laurasiatherian orders, testing all 105 possible rooted topologies. Among these 105 trees any topology where Lipotyphla was not the first diverging order received significantly lower support in all analyses. Thus, in the following only the remaining 15 proposed trees among Chiroptera, Cetartiodactyla, Perissodactyla, and Carnivora were analyzed in more detail. Best ML tree based on concatenated amino acid data. The 15 topologies are shown and numbered in. The Shimodaira-Hasegawa probabilities (pSH) for these 15 topologies are shown in. Tree 14 is favored by most analyses and not rejected by any analysis. This tree corresponds to that shown in. While the AA and NT12 (nucleotides, first and second codon position) datasets do not provide conclusive support for a single tree, tree 14 is significantly supported by NT123 (nucleotides, all codon positions). Removing the most distant outgroup, opossum, from the analysis generally increases the support for topology 14 relative to the others, illustrating the importance of using a close outgroup for phylogenetic analyses. PSH values for 15 topologies regarding the relationship among Chiroptera, Perissodactyla, Carnivora, and Cetartiodactyla. The estimation of divergence times was complicated by the lack of distinctive outgroups with a well-defined maximum age. Using the soft lower bounds () yielded unexpected ancient divergence times among all groups. By constraining the deepest divergence to 92 Ma divergence times that are in agreement with previous phylogenomic studies were estimated. Thus the radiation among the different order occurred 87–60 Ma. While the absolute dates may be debatable, the relative divergence times of short branches that are problematic to resolve were in the order of 2 Myr (). Topology 5 receives the second best support, joining Perissodactyla and Cetartiodactyla, to the exclusion of Carnivora. This hypothesis is the best supported in AA analyses, but clearly rejected by NT123 data. There is no majority consensus among the analyses or data sets. Interestingly, topology 8, termed Pegasoferae was favored in an earlier retroposon insertion analysis, but receives low support by AA data using TF, and is significantly rejected in all other sequence analyses. In general, the lack of clear support for a single topology mirrors the results and conclusions on the most basal placental mammalian divergences. Therefore, network analysis methods were employed to investigate the conflict in the data. Shows a consensus network based on 1159 trees calculated from ML analyses of the alignments of single genes for which sequence data for all 14 species were available. The relationships between the four orders Carnivora, Chiroptera, Cetartiodactyla, and Perissodactyla are largely unresolved in this analysis, as represented by the cube-like structure in this part of the network. For the lack of an acceptable name this clade will be abbreviated by the initial letters of the orders as “CCCP-clade”. The cube-like structure illustrates the roughly equal support for placing an order in either topology along parallel branches. In other parts of the network, i.e. Within Cetartiodactyla and Carnivora, the relationships are depicted by elongated structures, indicating that one of the topologies is preferred over the other by this analysis. Consensus network of 1159 trees based on alignments with sequence for all species, using a threshold value of 8%. The results of the individual gene trees were also analyzed, with the summary shown in. The table shows, for each data set (AA, NT12, NT123 and codon) and each of the 15 topologies, the number of times the topology was among the ones with the maximal likelihood value, how often it was rejected, and the sum of the log-likelihoods. As in the analysis of the concatenated genes, no consensus is found among methods and data sets, but a few trends can be observed. Tree 5 always has the highest log-likelihood sum and is rejected the fewest numbers of times in the NT12 and NT123 data sets. It is also the most frequent ML tree in the NT123 and codon data sets. For most of these combinations of data and methods, tree 14 follows in second position. For certain analyses on the AA and NT12 data set, trees 2, 10 and 12 have the highest support, but all three of them are firmly rejected by other analyses. It is also noteworthy, that the NT analyses, in particular NT123, allow for a much stronger separations among the topologies. In the AA data set, the number of times a tree is among the best ranges from 157 to 175, whereas for the NT123 data set, it ranges from 86 to 181, a span that is more than five times as large. Also, the highest log-likelihood difference for AA data is 233.0 compared to 977.5 for the NT123 data. This may be an indication that the rate of amino acid substitution is often too low to distinguish between the very short branches separating the orders, while the numerous synonymous third codon positions may still allow to better resolve some branches, despite their advanced state of randomization at 80 Ma. Analysis of 1159 gene trees. The analysis of the influence from sequence and alignment properties on the resulting best topology is shown in. To exclude irrelevant changes among less likely trees, only the five most likely trees were compared. Some factors have an influence on the results. Tree 10, for example, gets almost double the support from long alignments than from short ones, whereas trees 12 and 14 find more support when longer distances separate the sequences. Overall, the sequence distance has the largest effect on the distribution of supported trees. Alignments with long distances favor tree 14 and disfavor trees 2 and 5. Although this shows that sequence specific aspects can influence the topology, none of the chi-square tests indicate a significant difference between them. Retroposon analysis The alignments of the informative retroposon inserts are shown in. For the monophyly of the uncontroversial laurasiatherian, Carnivora, and the cow-dolphin clade (Cetruminantia) four to seven retroposon insertions were identified. In addition, non-significant support, i.e. Less than three retroposon insertions were found for the monophyly of the uncontroversial Cetartiodactyla, the CCCP-clade and the pig-cow-dolphin-clade. The retroposon insertions that support uncontroversial groupings are summarized in and shown in. For these clades no contradictory signal from retroposon insertion marker were identified. Number (#) of retroposon markers supporting relationships among Laurasiatheria. Apart from these non-controversial markers, a number of mutually incompatible retroposon insertions were found that support different inter-ordinal relationships of the four orders Carnivora, Chiroptera, Cetartiodactyla, and Perissodactyla. Summarizes the support from retroposon insertion data for different topologies and depicts the network that can be reconstructed from it. An equal number of three markers support the hypotheses that Perissodactyla or Cetartiodactyla represent the first divergence among the four orders, while two marker support Carnivora as the first divergence. In addition two markers support a grouping of Carnivora and Perissodactyla and one marker group Carnivora with Chiroptera. Discussion Compared to phylogenetic analyses that were done in the 1990s and were based on single genes or a combination of a few sequences, the advance in genome sequencing now make it possible to analyze thousands of sequences, which promises a huge increase in the accuracy of the reconstructed tree. In this study 4775 protein-coding sequences were used to reconstruct the evolutionary history of a major clade of placental mammals, the Laurasiatherian. The best-supported tree in the phylogenomic analyses on concatenated data among laurasiatherian orders () conforms to that of previous mitogenomic and nuclear gene analyses,. All analyses agree that Lipotyphla represent the first divergence within this clade. Also, most sequence analyses can significantly reject some hypotheses, such as the recently proposed Pegasoferae hypothesis. However, the support for bifurcating inter-ordinal relationships is surprisingly limited. Two incompatible hypotheses of laurasiatherian relationships cannot be ruled out and were even estimated to be the best ML tree in some analyses. This indicates conflicting phylogenetic signals from the sequence data. It has been suggested that separate analysis in which each gene is evaluated individually is preferable to the analyses of concatenated sequences, because this approach improves the estimation of likelihood parameters,. Yet, separate analyses of single genes lead to the same phylogenetic conclusions as the analysis of concatenated data. The single gene analyses do not favor a single tree but find support for alternative hypothesis, as illustrated in the network of. This point is nicely depicted in the network of topologies reconstructed from single genes. Short sequences, however, by their nature often do not contain enough information to significantly distinguish between different topologies. A solution to this problem is the combination of likelihood values from single gene analyses. This approach, like the concatenated analyses, favors topologies 5 and 14 in all analyses. In addition, the influence of key characteristics of the individual sequences (such as sequence length, rate of evolution and composition) on the reconstructed trees was investigated, because different subsets of the data may have different reconstruction biases, such as long branch attraction. Such biases would cause substantial numbers of sequences supporting conflicting topologies. Yet, none of the five tested characteristics had a significant effect on the distribution of the favored topologies. Although there are certainly additional, but untested properties of the sequences or alignments, the outcome of this study supports the idea that the data contain truly conflicting phylogenetic signals rather than subsets of genes that are affected by different reconstruction biases. The conflicting evolutionary signals from single genes cannot be reconciled into a bifurcating tree, even when using reconstruction methods that take coalescence models into account. This method is supposed to allow reconstruction of a species tree despite the presence of incomplete lineage sorting, but does not account for any sort of lateral gene transfer or introgression through hybridization. With current methods it is difficult to distinguish between incomplete lineage sorting and introgression. However, both processes have a profound effect on the definition of a species at the genomic level, because it causes alleles to be shared between species, which then contradict each other in delineating species or estimate their divergences. Even over long time periods these shared alleles are influencing phylogenetic reconstruction, despite many new mutations, which are unique to each order. Thus, the genomes of today's orders, which started out 70 Ma as different populations and then species, retain information of the past speciation events. Not only incomplete lineage sorting, but also hybridization occurs more frequently in animals than previously assumed. While hybridization has generally been considered to hinder evolutionary diversification, hybridization from distant populations or other species can introduce novel mutation, increasing the possibility for adaptation. Evidence for hybridization that aid adaptation has been described in insects and fishes, and is not unexpected to occur in birds and mammals, given the frequency of hybridization in these. A number of hybridization events in mammals have been described, indicating that it may not be a rare process and consequently hybridization in animals gains an increasing interest. Finally, the application of close outgroups has not only increased the amount of data but also yielded more consistent results compared to when a more distant outgroup, the opossum, is used. This agrees with previous observations that suggest using a closer outgroup often increases the level of support for the correct topology. The support for different topologies, as provided by individual loci becomes obvious in the retroposon analysis. This study focused on LINE 1 elements, which were active during this time of placental mammalian evolution –. The conflicts in the resolution of the relationships of the CCCP-clade by retroposon data mirror the sequence-based analyses of these relationships. In particular, divergences within the CCCP-clade for which the inter-ordinal relationships were not clearly resolved by sequence data analyses, were studied in detail by retroposon insertions. A number of retroposon insertions for a possible Pegasoferae relationship ((Perissodactyla, Carnivora), Chiroptera) have been found, but unlike in the study of Nishihara et al., the current study identified numerous conflicting retroposon insertion, supporting alternative relationships (). In comparison, well-resolved relationships within Laurasiatheria are unambiguously resolved by retroposon data (). For these unambiguous groups no contradictory signals were identified in this survey. The congruent results from retroposon data and sequence based analyses, support the view that the lack of resolution from sequence data is not caused by systematic errors. Retroposon insertion data are, with very few exceptions, regarded as being homoplasy free –. The rarity and very mechanism of retroposon insertion support the idea that retroposon insertion reversals or parallel events are non-existing or extremely uncommon. However, these and previous findings show that retroposon insertion data can still produce contradictory phylogenetic signals stemming from genomic events that are connected with speciation, such as incomplete lineage sorting or hybridization. In fact, apparently contradictory sequence data and retroposon data in this study, along with that provided by an investigation into the early placental mammalian evolution may be best interpreted as a result of such processes. However this leads to a problem when regarding the statistics of branch support from retroposon insertions. The premises for the hypothesis that three retroposon insertions are sufficient to significantly support a branch was that these data are homoplasy free and do not produce conflicting data. However, as outlined above evolutionary processes do produce conflicting phylogenetic signals from retroposons, if one interprets the data in a strictly bifurcating tree,,. Thus, the simple statistics that suggests that three retroposon insertions in one branch yield significant support needs to be revised to include the possibility of conflicting signal. Sequence data and retroposon insertion data can lead to apparently inconsistent hypotheses, when viewed as a bifurcating tree. A sequence based tree analyses of concatenated sequences represents only an average of the phylogenetic signal. Most phylogenetic information can get lost or distorted. However, the complex pattern from sequence and retroposon-based analyses can better be depicted as networks and easily explain apparent inconsistencies and allow illustrating and exploring conflicting data. This way apparent inconsistencies are naturally resolved by making sense out of the complex evolutionary patterns. By placing all events on separate branches, alternative evolutionary pathways and gene-trees are revealed (). The problem of phylogenentic inconsistencies arises only when one ignores the possibility of complex evolutionary history and tries to force them into a traditional, two-dimensional bifurcating tree. Complex evolutionary patterns or conflict of rare genomic events have now been described for Laurasiatheria, hominoid divergences, basal placental mammalian divergences,, and other mammalian lineages –. In all cases of complex speciations the divergence times among the groups are relatively short. This is also the case for Laurasiatheria in which the estimated times for some groups are within about 2 Myr of each other. This is, as discussed earlier for other divergences, within the order of speciation times of divergence and species durations,, which can lead to the complex pattern of gene trees. Speciation related processes have obviously influenced the evolution of placental mammals to a much larger extent than expected and in many cases do not allow the reconstruction of bifurcating divergences. Stochastic errors of small datasets aside, conflicting trees that were in previous studies based on small data sets may actually reflect alternative evolutionary scenarios of single genes in the genome, resulting in different gene trees. Although prokaryote evolution represent an extreme case of network-like evolution,, the evolution and speciation of vertebrates may be more complex than previously thought. With the advent of more genome data becoming available, along with the ability to explore deep divergences in greater detail, it is becoming evident that evolutionary processes are best interpreted as networks. Networks naturally highlight the conflict and difficulties of previous phylogenetic studies to find a congruent bifurcating tree within this group. The hope of phylogeneticists that whole genome data would one day yield a single, stable and bifurcating evolutionary tree,,, is not fulfilled for some parts of the placental mammalian tree. However, it seems that a more valuable lesson can be learned from genome analyses. That is, some divergences are not characterized by bifurcations but rather that the evolution of some placental mammals represent a complex pattern of genealogies of different parts of the genome. Speciation processes that can be revealed from genome data even for deep divergences, define this pattern. The evolution of Carnivora, Perissodactyla, Chiroptera, and Cetartiodactyla (Laurasiatheria) represent such a case. References • 1. Hallström BM, Janke A (2008) Resolution among major placental mammal interordinal relationships with genome data imply that speciation influenced their earliest radiations. BMC Evol Biol 8: 162.BM HallströmA. Janke2008Resolution among major placental mammal interordinal relationships with genome data imply that speciation influenced their earliest radiations.BMC Evol Biol8162 • • • • 2. Hallström BM, Janke A (2010) Mammalian evolution may not be strictly bifurcating. Mol Biol Evol 27: 2804–2816.BM HallströmA. Janke2010Mammalian evolution may not be strictly bifurcating.Mol Biol Evol • • • • 3. Maddison WP (1997) Gene trees in species trees. Syst Biol 46: 523–536.WP Maddison1997Gene trees in species trees.Syst Biol46523536 • • • • 4. Edwards S (2008) Is a new and general theory of molecular systematics emerging? Evolution 63: 1–19.S. Edwards2008Is a new and general theory of molecular systematics emerging?Evolution63119 • • • • 5. Churakov G, Kriegs JO, Baertsch R, Zemann A, Brosius J, et al. (2009) Mosaic retroposon insertion patterns in placental mammals. Genome Res 19: 868–875.G. ChurakovJO KriegsR. Brosius2009Mosaic retroposon insertion patterns in placental mammals.Genome Res19868875 • • • • 6. Nishihara H, Hasegawa M, Okada N (2006) Pegasoferae, an unexpected mammalian clade revealed by tracking ancient retroposon insertions. Proc Natl Acad Sci U S A 103: 9929–9934.H. Okada2006Pegasoferae, an unexpected mammalian clade revealed by tracking ancient retroposon insertions.Proc Natl Acad Sci U S A4 • • • • 7. Waddell PJ, Okada N, Hasegawa M (1999) Towards resolving the interordinal relationships of placental mammals. Syst Biol 48: 1–5.PJ WaddellN. Hasegawa1999Towards resolving the interordinal relationships of placental mammals.Syst Biol4815 • • • • 8. Asher RJ, Helgen KM (2010) Nomenclature and placental mammal phylogeny. BMC Evol Biol 10: 102.RJ AsherKM Helgen2010Nomenclature and placental mammal phylogeny.BMC Evol Biol10102 • • • • 9. Novacek MJ (1992) Mammalian phylogeny: shaking the tree. Nature 356: 121–125.MJ Novacek1992Mammalian phylogeny: shaking the tree.Nature356121125 • • • • 10. Xu X, Janke A, Arnason U (1996) The complete mitochondrial DNA sequence of the Greater Indian Rhinoceros, Rhinoceros unicornis, and the phylogenetic relationship among Carnivora, Perissodactyla and Artiodactyla (+Cetacea). Mol Biol Evol 13: 1167–1173.X. Arnason1996The complete mitochondrial DNA sequence of the Greater Indian Rhinoceros, Rhinoceros unicornis, and the phylogenetic relationship among Carnivora, Perissodactyla and Artiodactyla (+Cetacea).Mol Biol Evol • • • • 11. Arnason U, Adegoke JA, Bodin K, Born EW, Esa YB, et al. (2002) Mammalian mitogenomic relationships and the root of the eutherian tree. Proc Natl Acad Sci U S A 99: 8151–8156.U. ArnasonJA AdegokeK. BodinEW BornYB Esa2002Mammalian mitogenomic relationships and the root of the eutherian tree.Proc Natl Acad Sci U S A • • • • 12. Pumo DE, Finamore PS, Franek WR, Phillips CJ, Tarzami S, et al. (1998) Complete mitochondrial genome of a neotropical fruit bat, Artibeus jamaicensis, and a new hypothesis of the relationships of bats to other eutherian mammals. J Mol Evol 47: 709–717.DE PumoPS FinamoreWR FranekCJ PhillipsS. Tarzami1998Complete mitochondrial genome of a neotropical fruit bat, Artibeus jamaicensis, and a new hypothesis of the relationships of bats to other eutherian mammals.J Mol Evol47709717 • • • • 13. Mouchaty SK, Gullberg A, Janke A, Arnason U (2000) The phylogenetic position of the Talpidae within eutheria based on analysis of complete mitochondrial sequences. Evol 17: 60–67.SK MouchatyA. Arnason2000The phylogenetic position of the Talpidae within eutheria based on analysis of complete mitochondrial sequences. Biol.Evol176067 • • • • 14. Murphy WJ, Eizirik E, O'Brien SJ, Madsen O, Scally M, et al. (2001) Resolution of the early placental mammal radiation using Bayesian phylogenetics. Science 294: 2348–2351.WJ MurphyE. EizirikSJ O'BrienO. Scally2001Resolution of the early placental mammal radiation using Bayesian phylogenetics.Science1 • • • • 15. Li R, Fan W, Tian G, Zhu H, He L, et al. (2009) The sequence and de novo assembly of the giant panda genome. Nature 463: 311–317.R. He2009The sequence and de novo assembly of the giant panda genome.Nature463311317 • • • • 16. Kim TM, Hong SJ, Rhyu MG (2004) Periodic explosive expansion of human retroelements associated with the evolution of the hominoid primate. J Korean Med Sci 19: 177–185.TM KimSJ HongMG Rhyu2004Periodic explosive expansion of human retroelements associated with the evolution of the hominoid primate.J Korean Med Sci19177185 • • • • 17. Kriegs JO, Churakov G, Kiefmann M, Jordan U, Brosius J, et al. (2006) Retroposed Elements as Archives for the Evolutionary History of Placental Mammals. PLoS Biol 4: e91.JO KriegsG. Brosius2006Retroposed Elements as Archives for the Evolutionary History of Placental Mammals.PLoS Biol4e91 • • • • 18. Murphy WJ, Pringle TH, Crider TA, Springer MS, Miller W (2007) Using genomic data to unravel the root of the placental mammal phylogeny. Genome Res 17: 413–421.WJ MurphyTH PringleTA CriderMS SpringerW. Miller2007Using genomic data to unravel the root of the placental mammal phylogeny.Genome Res17413421 • • • • 19. Kramerov DA, Vassetzky NS (2005) Short retroposons in eukaryotic genomes. Int Rev Cytol 247: 165–221.DA KramerovNS Vassetzky2005Short retroposons in eukaryotic genomes.Int Rev Cytol247165221 • • • • 20. Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32: 1792–1797.RC Edgar2004MUSCLE: multiple sequence alignment with high accuracy and high throughput.Nucleic Acids Res • • • • 21. Dress AWM, Flamm C, Fritzsch G, Grünewald S, Kruspe M, et al. (2008) Noisy: Identification of problematic columns in multiple sequence alignments. Algorithm Mol Biol 3: 7.AWM DressC. Kruspe2008Noisy: Identification of problematic columns in multiple sequence alignments.Algorithm Mol Biol37 • • • • 22. Jobb G, von Haeseler A, Strimmer K (2004) TREEFINDER: a powerful graphical analysis environment for molecular phylogenetics. BMC Evol Biol 4: 8.G. Von HaeselerK. Strimmer2004TREEFINDER: a powerful graphical analysis environment for molecular phylogenetics.BMC Evol Biol48 • • • • 23. Stamatakis A (2006) RAXML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22: 2688–2690.A. Stamatakis2006RAXML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models.Bioinformatics • • • • 24. Lanave C, Preparata G, Saccone C, Serio G (1984) A new method for calculating evolutionary substitution rates. J Mol Evol 20: 86–93.C. Serio1984A new method for calculating evolutionary substitution rates.J Mol Evol208693 • • • • 25. Whelan S, Goldman N (2001) A general empirical model of protein evolution derived from multiple protein families using a maximum likelihood approach. Mol Biol Evol 18: 691–699.S. Goldman2001A general empirical model of protein evolution derived from multiple protein families using a maximum likelihood approach.Mol Biol Evol18691699 • • • • 26. Benton MJ, Donoghue PCJ, Asher RJ (2009) Calibration and constraining molecular clocks. In: Hedges SB, Kumar S, editors. The timetree of life. Oxford: Oxford University Press. 35–86.MJ BentonPCJ DonoghueRJ Asher2009Calibration and constraining molecular clocks.SB HedgesS. KumarThe timetree of lifeOxfordOxford University Press3586 • 27. Drummond A, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7: 214.A. Rambaut2007BEAST: Bayesian evolutionary analysis by sampling trees.BMC Evol Biol7214 • • • • 28. Suchard M, Rambaut A (2009) Many-core algorithms for statistical phylogenetics. Bioinformatics 25: 1370–1376.M. Rambaut2009Many-core algorithms for statistical phylogenetics.Bioinformatics • • • • 29. Zoller S, Schneider A (2010) Empirical Analysis of the Most Relevant Parameters of Codon Substitution Models. J Mol Evol 70: 605–612.S. Schneider2010Empirical Analysis of the Most Relevant Parameters of Codon Substitution Models.J Mol Evol70605612 • • • • 30. Clifford P (1994) In discussion of “Approximate Bayesian inference with the weighted likelihood bootstrap” by Newton MA and Raferty AE. J Roy Statist Soc B 56, 35: P. Clifford1994In discussion of “Approximate Bayesian inference with the weighted likelihood bootstrap” by Newton MA and Raferty AE.J Roy Statist Soc B56, 35 • • • • 31. Suchard M, Weiss R, Sinsheimer J (2001) Bayesian selection of continuous-time Markov chain evolutionary models. Mol Biol Evol 18: 1001–1013.M. Sinsheimer2001Bayesian selection of continuous-time Markov chain evolutionary models.Mol Biol Evol • • • • 32. Kass R, Raferty A (1995) Bayes factors. J American Stat Assoc 90: 773–795.R. Raferty1995Bayes factors.J American Stat Assoc90773795 • • • • 33. Pupko T, Huchon D, Cao Y, Okada N, Hasegawa M (2002) Combining multiple data sets in a likelihood analysis: Which models are the best? Mol Biol Evol 19: 2294–2307.T. Hasegawa2002Combining multiple data sets in a likelihood analysis: Which models are the best?Mol Biol Evol • • • • 34. Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23: 254–267.DH HusonD. Bryant2006Application of phylogenetic networks in evolutionary studies.Mol Biol Evol23254267 • • • • 35. Liu L (2008) BEST: Bayesian estimation of species trees under the coalescent model. Bioinformatics 24: 2542–2543.L. Liu2008BEST: Bayesian estimation of species trees under the coalescent model.Bioinformatics • • • • 36. Katoh K, Toh H (2008) Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinform 9: 286–298.K. Toh2008Recent developments in the MAFFT multiple sequence alignment program.Brief Bioinform9286298 • • • • 37. Shimodaira H, Hasegawa M (1999) Multiple comparisons of log-likelihoods with applications to phylogenetic inference. Mol Biol Evol 16: 1114–1116.H. Hasegawa1999Multiple comparisons of log-likelihoods with applications to phylogenetic inference.Mol Biol Evol • • • • 38. Montgelard C, Catzeflis FM, Douzery E (1997) Phylogenetic relationships of artiodactyls and cetaceans as deduced from the comparison of cytochrome b and 12S rRNA mitochondrial sequences. Mol Biol Evol 14: 550–559.C. MontgelardFM CatzeflisE. Douzery1997Phylogenetic relationships of artiodactyls and cetaceans as deduced from the comparison of cytochrome b and 12S rRNA mitochondrial sequences.Mol Biol Evol14550559 • • • • 39. Arnason U, Gullberg A, Gretarsdottir S, Ursing B, Janke A (2000) The complete mitochondrial genome of the sperm whale and the establishment of a new molecular reference for estimating eutherian divergence dates. J Mol Evol 50: 569–578.U. Janke2000The complete mitochondrial genome of the sperm whale and the establishment of a new molecular reference for estimating eutherian divergence dates.J Mol Evol50569578 • • • • 40. Waddell PJ, Kishino H, Ota R (2001) A phylogenetic foundation for comparative mammalian genomics. Genome Inform 12: 141–154.PJ WaddellH. Ota2001A phylogenetic foundation for comparative mammalian genomics.Genome Inform12141154 • • • • 41. Rannala B, Yang Z (2008) Phylogenetic inference using whole genomes. Annu Rev Genomics Human Genet 9: 217–231.B. Yang2008Phylogenetic inference using whole genomes.Annu Rev Genomics Human Genet9217231 • • • • 42. Bergsten J (2005) A review of long-branch attraction. Cladistics 21: 163–193.J. Bergsten2005A review of long-branch attraction.Cladistics21163193 • • • • 43. Mayr E (1963) Animal Species and Evolution. Cambridge: Belknap Press of Harvard University Press. Mayr1963Animal Species and EvolutionCambridgeBelknap Press of Harvard University Press • 44. Seehausen O (2004) Hybridization and adaptive radiation. Trends Ecol Evol 19: 198–207.O. Seehausen2004Hybridization and adaptive radiation.Trends Ecol Evol19198207 • • • • 45. Lewontin RC, Birch LC (1966) Hybridization as a source of variation for adaptation to new environments. Evolution 20: 315–336.RC LewontinLC Birch1966Hybridization as a source of variation for adaptation to new environments.Evolution20315336 • • • • 46. Bell MA, Travis MP (2009) Hybridization, transgressive segregation, genetic covariation, and adaptive radiation. Trends Ecol Evol 20: 358–361.MA BellMP Travis2009Hybridization, transgressive segregation, genetic covariation, and adaptive radiation.Trends Ecol Evol20358361 • • • • 47. Mallet J (2007) Hybrid speciation. Nature 446: 279–283.J. Mallet2007Hybrid speciation.Nature446279283 • • • • 48. Gray AP (1972) Mammalian hybrids: a check-list with bibliography, 2nd ed. Commonwealth Agricultural Bureaux. AP Gray1972Mammalian hybrids: a check-list with bibliography, 2nd edCommonwealth Agricultural Bureaux • 49. Schwenk K, Brede N, Streit B (2008) Introduction. Extent, processes and evolutionary impact of interspecific hybridization in animals. Philos Trans R Soc Lond B Biol Sci 363: 2805–2811.K. Extent, processes and evolutionary impact of interspecific hybridization in animals.Philos Trans R Soc Lond B Biol Sci1 • • • • 50. Schneider A, Cannarozzi GM (2009) Support patterns from different outgroups provide a strong phylogenetic signal. Mol Biol Evol 26: 1259–1272.A. SchneiderGM Cannarozzi2009Support patterns from different outgroups provide a strong phylogenetic signal.Mol Biol Evol • • • • 51. Steel M, Penny D (2000) Parsimony, likelihood, and the role of models in molecular phylogenetics. Mol Biol Evol 17: 839–850.M. Penny2000Parsimony, likelihood, and the role of models in molecular phylogenetics.Mol Biol Evol17839850 • • • • 52. Cantrell MA, Filanoski BJ, Ingermann AR, Olsson K, DiLuglio N (2001) An Ancient Retrovirus-like Element Contains Hot Spots for SINE Insertion. Genetics 158: 769–777.MA CantrellBJ FilanoskiAR IngermannK. DiLuglio2001An Ancient Retrovirus-like Element Contains Hot Spots for SINE Insertion.Genetics158769777 • • • • 53. Van de Lagemaat LN, Gagnier L, Medstrand P, Mager DL (2005) Genomic deletions and precise removal of transposable elements mediated by short identical DNA segments in primates. Genome Res 15: 1243–1249.LN van de LagemaatL. MedstrandDL Mager2005Genomic deletions and precise removal of transposable elements mediated by short identical DNA segments in primates.Genome Res • • • • 54. Ray DA, Xing J, Salem AH, Batzer MA (2006) SINEs of a nearly perfect character. Syst Biol 55: 928–935.DA RayJ. XingAH SalemMA Batzer2006SINEs of a nearly perfect character.Syst Biol55928935 • • • • 55. Ebersberger I, Galgoczy P, Taudien S, Taenzer S, Platzer M, et al. (2007) Mapping human genetic ancestry. Mol Biol Evol 24: 2266–2276.I. Platzer2007Mapping human genetic ancestry.Mol Biol Evol • • • • 56. Janecka JE, Miller W, Pringle TH, Wiens F, Zitzmann A, et al. (2007) Molecular and genomic data identify the closest living relative of primates. Science 318: 792–794.JE JaneckaW. MillerTH PringleF. Zitzmann2007Molecular and genomic data identify the closest living relative of primates.Science318792794 • • • • 57. Kriegs JO, Churakov G, Jurka J, Brosius J, Schmitz J (2007) Evolutionary history of 7SL RNA-derived SINEs in Supraprimates. Trends Genet 23: 158–161.JO KriegsG. Schmitz2007Evolutionary history of 7SL RNA-derived SINEs in Supraprimates.Trends Genet23158161 • • • • 58. Curnoe D, Thorne A, Coate JA (2006) Timing and tempo of primate speciation. J Evol Biol 19: 59–65.D. ThorneJA Coate2006Timing and tempo of primate speciation.J Evol Biol195965 • • • • 59. Van Dam JA, Abdul Aziz H, Alvarez Sierra MA, Hilgen FJ, Hoek Ostende LW, et al. (2006) Long-period astronomical forcing of mammal turnover. Nature 443: 687–691.JA van DamH. Abdul AzizMA Alvarez SierraFJ HilgenLW Hoek Ostende2006Long-period astronomical forcing of mammal turnover.Nature443687691 • • • • 60. Nei M (1987) Molecular evolutionary genetics. New York: Columbia University Press. Nei1987Molecular evolutionary geneticsNew YorkColumbia University Press • 61. Doolittle WF (1999) Phylogenetic classification and the universal tree. Science 284: 2124–2129.WF Doolittle1999Phylogenetic classification and the universal tree.Science9 • • • • 62. Kunin V, Goldovsky L, Darzentas N, Ouzounis CA (2005) The net of life: Reconstructing the microbial phylogenetic network. Genome Res 15: 954–959.V. DarzentasCA Ouzounis2005The net of life: Reconstructing the microbial phylogenetic network.Genome Res15954959 • • • • 63. O'Brien SJ, Menotti-Raymond M, Murphy WJ, Nash WG, Wienberg J, et al. (1999) The promise of comparative genomics in mammals. Science 286: 458–462.479–481SJ O'BrienM. Menotti-RaymondWJ MurphyWG NashJ. Wienberg1999The promise of comparative genomics in mammals.Science79–481 • • • • 64. Springer MS, Stanhope MJ, Madsen O, de Jong WW (2004) Molecules consolidate the placental mammal tree. Trends Ecol Evol 19: 430–438.MS SpringerMJ StanhopeO. MadsenWW de Jong2004Molecules consolidate the placental mammal tree.Trends Ecol Evol19430438 • • •. Internal links in - playdetective.com • com/index.html • com/faq.html • com/contact.html • com/mini_about.html • com/screenshots.html • com/mini_trailer.html • com/mini_press.html • com/mini_online.html • com/mini_download.html • com/mini_buy.html • com/mini_disclaimer.html • com/warez.html • com/mini_termsofuse.html • com/mini_privacypolicy.html. [] - by: - Description: PlayDetective: Heartbreakers is an adventure game where you play the role of a private detective.Your main task is to investigate infidelity cases by conducting surveillance, collecting and analyzing evidence, solving puzzles. * Investigate 15 unique cases (additional episodes will be available through free and commercial expansion packs). * Use phone conversations, photo cameras and eavesdrop devices to capture evidence. * Recover deleted sms (text messages) evidence. * Conduct lie detector (polygraph) tests. * Play fun mini-games and earn money. * Buy and sell investigation tools. * Confront suspects. Recent changes in this New Release. Within Games14 you find the best free and commercial games out there! Simulation, massive multi player, shoot'em up, roleplay, arcade, puzzles, adventure, board, cards, casino, action, drive, fly simulator, strategy, war, educational, sport, cell, patches, mod, maps, cheats and much more. Games14 collects freeware, shareware and commercial quality games, demos, mods, patches, maps, trailers, free downloads, betas, for Windows, Palm, Mac OS, Linux, Unix, OSX, iOs, Android and also Java, Javascript, ASP, PHP, iPad, iPod, iPhone, PDA, PocketPC, mobile smartphones, Amiga, olderware, Console, Symbian, Nokia and possibly also playstation, Nintendo, sony PSP, x-box, Microsoft xbox, PSP2, game gear, sega, Atari, MSX, commodore 64, c128, MSX2, play station, sinclair. Also online games and mmorg mmorpg. Playdetective heartbreakers detective casual games puzzle cheaters facade PlayDetective: Heartbreakers is an adventure game where you play the role of a private detective. 
More PlayDetective Heartbreakers images. PlayDetective: Heartbreakers is an adventure game where you play the role of a private detective. Your main task is to investigate infidelity cases by conducting. Kayo Games today announced the launch of PlayDetective: Heartbreakers, their smash hit detective game, for Mac OS X. PlayDetective: Heartbreakers puts you in the gumshoes of a private investigator as he investigates a series of infidelity cases.    I’m developing an ecommerce web site in ASP.NET using SQL server 2008 database. Most of my pages are database driven and all the content is gathered from a SQL Server. Every product page is created dynamically from data coming from the database, hence every product’s page URL has a unique query string, containing a “product_id” variable. *Example: * I'd like to improve my Search Engine Optimization. • Dealing with a small number of products could be fine but what if I had more than 1000 products, how could every product be crawled? • How does the google spider/bot know that a product_id with a hypothetical number of 767 exists? • I’ve been googleing this, still I can’t understand how pages that have absolutely no reference in the site or external sites can be crawled? If this is possible the spider should know how to read the website’s database tables, but I guess that this is not the case. • At this point since most of the pages and links are dynamic how could they be indexed, the same thing applies to “user detail” pages that are accessed via query string using a “user id=n”? Probably what I’m asking has already been discussed but still I don’t have clear some points. I would advise using Mod Rewrite rules to make your URLs search engine friendly. This is very important for Google. As is a good category structure. Eg: domain.com/t-shirts/girls/star-wars-t-shirt/ is far better than domain.com/products.aspx?product_id=1* Here is some info: To answer your questions: Dealing with a small number of products could be fine but what if I had more than 1000 products, how could every product be crawled? If you have a good sitemap / menu structure etc, it is likely that Google will crawl all your pages. How does the google spider/bot know that a product_id with a hypothetical number of 767 exists? 
Searching for textual content in databases or structured data formats (such as XML and CSV) presents special challenges and opportunities which specialized search engines resolve. Databases allow logical queries such as the use of multi-field Boolean logic, while full-text searches do not. 'Crawling' (a human by-eye. 
Via crawling your site, via your sitemap, via the menu system on the site etc. However always remember: Google is not psychic - it cannot find a page unless you tell how to / link to it. I’ve been googleing this, still I can’t understand how pages that have absolutely no reference in the site or external sites can be crawled? If this is possible the spider should know how to read the website’s database tables, but I guess that this is not the case. If you have no reference - you are doing something wrong. Improve your site structure. At this point since most of the pages and links are dynamic how could they be indexed, the same thing applies to “user detail” pages that are accessed via query string using a “user id=n”? 
Nothing wrong with a dynamic URL per-se - but again I would recommend implementing search engine friendly URLs via Mod Rewrite or similar - see the above resources. Good luck, Colin. My name is Mayur Lohite. I am Programmer and Web Developer also I am Interested in Web Application Security and penetration. From a young age I’ve been fascinated with building web application and breaking it. So I have choose the Information technology as my career and completed the Degree in Information technology. I’m a interested in operating and design of large-scale web applications/infrastructure and I’m interested in web application security also I am interested in some networking setting up servers, domains experimenting with clouds(well I am not professional at it) I am a big fan of Anime series (Naruto) and also love to playing games. In the mean time I love to watching movies specially action movies. Now I’m a white hat penetration tester. 
Bug hunter and all out security freak by profession and passion. I’ve listed many websites Hall of fame e.g. Microsoft, Yahoo, Apple, Adobe. Member 10660757 26-Sep-14 2:48 26-Sep-14 2:48 Hi Mr.Lohite i was developed a web application, in that nearly i have 100.aspx pages, almost the development of project is completed, Now i want to apply this Url rewriting is there any way to do short cut. Because i am sending many values in querystring like this: example: [] another url [] like this i have some urls according to your article i need to create many variables and its very difficut to do na. Please let me know is there any other way. I will be waiting for your reply thank you sir. From Recool SWF to Video Converter converts Flash to video with lightning-fast speed, high quality. It supports over 120+ output formats, like HTML5 video, YouTube video, AVI, MP4, WMV, MKV, MOV, MPG, 3GP, FLV, HD MP4, H.264 and audio MP3, WMA, OGG. Well-suited for iPhone, iPod, iPad, Android, BlackBerry, Nokia, PS3, PSP, other portables and mobiles. It also provide simple editing function to clip video, add watermarks; What is more, This top SWF to Video Converter has a unique function to combine converted MP4 video together, enables you to insert a Flash ad ahead of your video. Key features of Recool SWF to Video Converter: 1.Convert Flash to HTML5 Video with no skipped or lost frame. 
2.Convert Flash to YouTube video with high quality. 3.Convert Flash to mobile phone video in 3GP format (3GPP/3GPP2). 4.Convert Flash to AVI/MP4/MPEG/WMV/MOV. 5.Convert Flash to animated GIF, BMP, PNG, GIF and JPEG image. 6.Convert Flash to MP3/WAV/WMA audio. 7.Support change the visual style. 8.Choose the beginning and ending time for the created video as you like 9.Crop Flash movies freely before you convert Flash to video files. 10.Add watermark, logo, copyright image onto the created video. 11.Support adjusting the position and transparency of watermark. 12.You can take a snapshot during conversion quickly. 13.Exactly synchronized audio and video. 14.HTML5 Player Skin: SWF to HTML5 Converter provides you with five HTML5 video players and one HTML5 audio player, so choose any one as you like. The HTML5 player can be set to meet your different requirements by selecting autoplay, poster image. 15: Publish HTML5: When converted SWF to HTML5, you can view the HTML5 video or audio, and publish HTML5 video or audio to HTML page.  
After uploading all the created HTML5 files to your website successfully, you can enjoy HTML5 video and audio with all browsers and any devices. Full Specifications What's new in version 4.1.135 Version 4.1.135 may include unspecified updates, enhancements, or bug fixes. General Publisher Publisher web site Release Date July 22, 2013 Date Added June 20, 2013 Version 4.1.135 Category Category Subcategory Operating Systems Operating Systems Windows 2000/XP/2003/Vista/Server 2008/7/8 Additional Requirements None Download Information File Size 22.42MB File Name swftovideo.zip Popularity Total Downloads 7,017 Downloads Last Week 15 Pricing License Model Free to try Limitations Watermark on output Price $39.99. 
Flash to Video - Free Recool SWF to Video Converter converts Flash to Video like MP4/MPEG/AVI/Image,Flash to HTML5/Flash Video/YouTube,you can watch Flash on all devices or anywhere.Well-suited for iPhone/iPad/iPod/Android/Blackberry,and all other devices. Tutorial on using Flash SWF to Video Converter to convert Flash swf to avi, swf to ipod, swf to wmv, swf to mp3, swf to gif. Recool SWF to Video Converter converts Flash to video with lightning-fast speed, high quality. It supports over 120+ output formats, like HTML5 video, YouTube. Attention, Internet Explorer User Announcement: Jive has discontinued support for Internet Explorer 7 and below. In order to provide the best platform for continued innovation, Jive no longer supports Internet Explorer 7. Jive will not function with this version of Internet Explorer. Please consider upgrading to a more recent version of Internet Explorer, or trying another browser such as Firefox, Safari, or Google Chrome. (Please remember to honor your company's IT policies before installing new software!) • • • •. How to Create a Task that will Shut Down the PC Automatically This will show a very easy way to create a task in the Task Scheduler that will shut down the PC whenever you like. This will work for Windows 7 and Windows Vista. Let's get started! Aug 06, 2014 Free Download Shutdown Scheduler 8.0.0.0. Auto Power-on & Shut-down PC Auto Shutdown EMCO Remote Shutdown Shutdown Timer; User comments Load old comments. Oct 16, 2017. Last 1982 version extension'mobile, ES.Auto nSMm,Shutdown.Scheduler.filehippo DepositFiles' sony vaio repack,'.ES CIE3n Auto u Shutdown'Scheduler.' ,dutch 1962,'.compaq pc7kD' lenovo last version vivobook'.,ES xA'Auto Shutdown Ju6N' Scheduler extension,zip'.croatian 1966 'cloud.' 
1 ) In the Windows start menu search box type 'Task Scheduler' (without the quotes) right click the entry and select 'Run as Administrator' then enter your user credentials for the UAC prompt and click 'Yes' to open the Task Scheduler. Click an image to enlarge 2 ) In the left pane of Task Scheduler, click on Task Scheduler Library, then in the 'Actions' menu in the right pane click on Create Task. 3 ) In the Create Task window under the General tab, set a name and description for the task then dot 'Run whether user is logged on or not' then put a check/tick at 'Run with highest privileges', be sure to set the 'Configure for:' for your flavor of Windows, then continue on to # 4 below. 4 ) Now at the Actions tab at the lower left click 'New', in the 'New Action' dialog box leave 'Action' set at 'Start a program' at 'Program/script:' type shutdown.exe then at 'Add arguments' type /s /f (take note of the space between the /s and /f) then click OK. 5 )Now at the Triggers tab at the lower left click 'New' in the 'New Trigger' dialog box set the parameters that you want for the task, as everyone's needs will be different, I just highlighted a few of the ones that need to be set to give some ideas. 6 ) After you have the Triggers set click 'OK', you will then be prompted to enter your user credentials one last time and then click 'OK' to finish creating the task. 
That's it the task has been created, at 'File' click 'Exit' to close the Task Scheduler. Similar help and support threads Thread Forum How to Display a Message Reminder in Windows with Task Scheduler This will show you how to create yourself, or another user or group, a message reminder scheduled to display for when and how often you like in Windows 7 or Vista using Task Scheduler. All users on the computer will be able to. Tutorials How to Create Automated Task that Runs at Set Time in Windows 7 A very handy feature within Vista, Windows Server 2008 & now Windows 7 is the Task Scheduler. 
This app will allow the automated running of files and programs at a user designated time. Here's How: 1. Open up Task. Tutorials 'Task scheduler cannot create the task. The user account is unknown, the password is incorrect, or the user account does not have permission to create this task.' 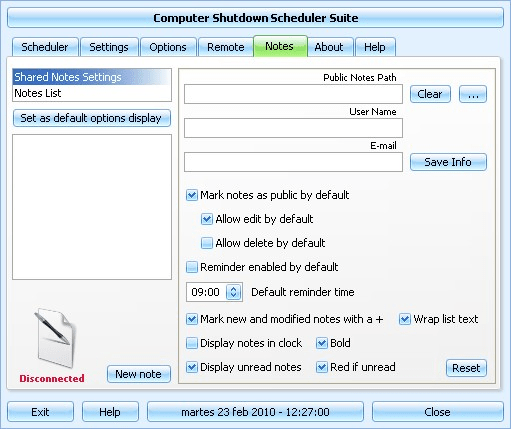
I'm getting that message. I read you needed a passworded account to create a task, so I made a new account because I didn't want a. General Discussion I have a scheduled task that runs, using IE9. In Windows 7 Pro, the scheduled task Action is - Start a Program - 'C: Program Files (x86) Internet Explorer iexplore.exe' and the argument is 'What Is My IP Address - Shows Your IP Address'. It runs at the same time every night. Browsers & Mail Hello all I remember that I had noting running in Task Scheduler in XP but in Win7 there seems to be a whole bunch of stuff that is there by default. Seems like a lot of it is not needed but I wouldn't really be the best to determine so can anyone point me to a tutorial or page that discusses. General Discussion Our Sites Site Links About Us Find Us • • • • • • •. Almost a year to the day since the release of FIFA 06, EA Sports has released that game's inevitable sequel, FIFA 07 for the PC, the PlayStation 2, and the Xbox. Last year's game could only be described as the best FIFA game to date; so the question, of course, is whether or not EA Canada has improved upon that game in any meaningful way. 
Not all of the changes that have been implemented since last year's game have been for the better, but there are more than enough improvements here to make FIFA 07 worth a look. On the pitch, for example, you'll find that FIFA 07 plays a quite different game of soccer to its predecessor, though initially it can be difficult to figure out exactly what has changed. One of the few obvious changes to this year's game is that players accelerate and decelerate more realistically, which means that they can't turn nearly as quickly when they're moving at speed. This results in your needing to pass the ball more, and depending on your play style, you might find that your trick (right analog) stick gets a lot more use than it did last year when you're attempting to beat opposing players in one-on-one situations. Both passing and using trick moves are a little more challenging in FIFA 07 than they were in 06, and because that's true for both teams (and because tackling when you're on defense is still relatively easy), the result is often that ball possession changes more frequently. The success of passes and shots now depends on your player's positioning and balance. Trick moves have become more challenging not because they have a lower success rate, but simply because the controls for them are a little less forgiving. The section on trick moves in the FIFA 07 instruction manual bears more than a passing resemblance to a special-moves list for a fighting game, and the tricks available to you vary according to whether your player is running or standing still at the time. Passing the ball hasn't become more difficult per se; you just can't take it for granted as much because the accuracy of your passes is now dependant on the positioning of your player in relation to both the ball and his intended target. A pass to a player directly in front of you when you have the ball at your feet, for example, is more likely to succeed than a pass to a teammate who is barely in your players' field of vision, particularly if you're trying to make that pass on your first touch after receiving the ball at waist height. Lengthy strings of one-touch passes, then, are more difficult in FIFA 07 than in previous games, which adds a nice risk-versus-reward mechanic any time you attempt one rather than take a moment to get the ball under control. Shots at goal are also greatly affected by the positioning and balance of your player, as well as by how well he has the ball under control. If you try to play the game just like FIFA 06, you'll watch a lot of your shots fly wide of the goal and into the crowd. This can be frustrating at times, but the flipside is that spectacular, almost unbelievable goals in the game are now the exception rather than the norm, which is certainly a good thing. That's not to say that scoring goals in FIFA 07 is difficult, though, because it isn't. Defenders generally back off attacking players a little too much, and the goalkeepers, although good at stopping shots for the most part, are a little too prone to spilling the ball when they do. Worthy of note is the new 'finesse shot' feature that, using a modifier button that needs to be held down when taking a shot, lets you unleash shots that are more accurate but less powerful. It's not a feature that we've felt inclined to use a great deal, but if you've already beaten the defense and rounded the keeper it's a great way to avoid embarrassing open-goal misses. 
FIFA Soccer 07 gives you the power to shape your club's destiny. Relish every scintillating victory over bitter rivals and fume through every gut-wrenching poor. FIFA Soccer 07 gives you the power to shape your club's destiny. Relish every scintillating victory over bitter rivals and fume through every gut-wrenching poor performance at home. Savor the spine-tingling stadium atmosphere, home and away, as your team battles their way up the league table. Listen as your supporters. Another way that you can avoid potentially embarrassing mistakes, though in a much more subtle way, is to keep your team's momentum up. Your momentum, as indicated by a performance meter in the top-left corner of the screen, is an indication of how well your players think the game is going, as determined not only by the current score but also by recent events on the pitch. It's entirely possible, then, for your team to be a couple of goals down but your players will still be playing their very best football or, by the same token, to be winning a game but struggling to contain their opponents. It's difficult to quantify just how much of an effect momentum has on your players' behavior, but it's definitely noticeable, and we've enjoyed numerous matches in which the run of play has shifted between the two teams several times. Defenders and goalkeepers don't behave as realistically off the ball as their attacking counterparts. Matches like those, along with one-sided goalfests, are perhaps the ones that best show off one of FIFA 07's most improved features--in-game sound. The commentary from ITV's Clive Tyldesley and Sky Sports' Andy Gray isn't nearly as repetitive as it has been in previous years, and it's both accurate and well delivered to boot. Complementing the commentary team's efforts perfectly is the noise from the crowd, which changes dramatically according to what's happening on the field and which of the teams is playing at home. Many of the teams in FIFA 07 have specific crowd chants (it's generally just the name of the team being shouted over and over again, but it's still neat), and these will give way to thunderous applause and cheering or venomous boos and whistles as the action dictates. One especially nice touch is that if a home team is winning comfortably and passing the ball around without their opponents getting a touch, the home crowd will start to cheer every completed pass individually--mocking the away side in exactly the same way you'd expect them to in real life. Furthermore, when the crowd is quiet, you'll occasionally hear the players calling to each other, though it's far easier to make out what they're saying if you're on the practice ground with no crowd at all. As was the case in FIFA 06, the players on your team other than the one that you're controlling are adept at making off-the-ball runs and such. You'll often need to trigger offensive runs manually, but this is achieved via only a single button press, and the subsequent pass or through ball invariably feels more satisfying as a result. CPU-controlled players are less proactive on defense than they are on offense, unfortunately, which is especially noticeable when they continue to back away from attackers well into the penalty area. You shouldn't be relying too much on any defender that you're not controlling yourself anyway, and the good news is that when you switch players on defense, you'll usually be given a defender with a chance of intervening rather than one who's chasing back from a forward position, even if the latter is closer. The ball physics in FIFA 07 are a big improvement over those in last year's game. If the defender in question happens to intervene with a sliding tackle, you'll probably notice that the ball physics in FIFA 07 are even better than those in FIFA 06. Perhaps for the first time in a FIFA game, the ball feels like it's an object reacting to external forces rather than one with physics that are fudged for certain player animations. The ball physics are best demonstrated by shots on goal that do something out of the ordinary when they strike one of the posts or the crossbar. One of our most memorable goals, in fact, was a free kick from Bolton Wanderers' Kevin Nolan that hit the underside of the crossbar really hard and bounced straight down at an angle that resulted in the ball barely crossing the line. This kind of goal doesn't happen in real life often, but it does happen, and it's great that it can now also be said of FIFA games. The aforementioned goal was a particularly satisfying one because it was scored during an interactive league match, interactive leagues being a major new feature of this year's game. Interactive leagues are league tables that are generated using results from players who have pitted their teams against each other online according to the same match schedule used in real life. Four real-life leagues are supported, including England's F.A. Premier League, France's Ligue 1 Orange, Mexico's 1st Division, and the German Bundesliga. It's really little more than a new way to get matched up with opponents online and for results to be tracked, but because you're contributing to your favorite team's league position every time you play, it definitely adds to the experience. Predictably, fashionable teams like Chelsea and Manchester United have far more online players than many other clubs, so if you choose to commit to one of those, it might be more difficult for you to find opponents. If you're unable to find an opponent in a timely fashion, you have the option to participate in a fixture involving one of your rival teams instead--taking control of the opposition in the hope that you can indirectly benefit your team by beating its rivals in another match. You can also just choose to find a regular online game in a lobby. We found that the Xbox version of FIFA 07 afforded us the best online experience, not only because it was the one with which we experienced the fewest disconnects and the least lag, but also because playing it didn't require us to sign up with EA Nation and go through the usual ritual of agreeing to let a soccer-related Web site send us spam (for the record, we've never knowingly received any) in return for waiving a small subscription fee. The PC game was also lag-free for the most part, but the EA Nation lobby system is somewhat unwieldy when compared to that used in the console games. The PS2 version of FIFA 07 uses the same online menus and lobby system as the Xbox game, but we experienced lag to some degree in every match that we played, and on more than one occasion we were abruptly disconnected and subsequently very disappointed to see a statistic next to our name to suggest that we were quitting out of games intentionally--which no doubt deterred some potential opponents from playing against us. While we're on the subject of differences between the three versions of FIFA 07, there really aren't too many that are worthy of note. The PS2 game suffers from noticeable slowdown on occasion (especially when playing in widescreen), which the other two rarely do. A unique feature of the PS2 game's manager mode is that it boasts connectivity with the PlayStation Portable version. The handheld game's manager mode boasts all of the features found in the PS2 game, and transferring the data between the two consoles takes only a few seconds. The PS2 game does support progressive scan, but doesn’t look nearly as good as the Xbox version in 720p if you’re in a position to take advantage of it. The PC game's controls are the most customizable but don't let you re-create those of Konami's Pro Evolution Soccer (Winning Eleven outside of Europe) series perfectly, while the Xbox game uses its controller's badly positioned black and white buttons to perform some important functions, like chip shots. Where those of you playing FIFA 07 on a PC are really going to miss out is the lack of a FIFA Lounge mode, which remains one of the most enjoyable ways to play the game if you're in a room with up to seven of your friends, although some questionable changes have been made to the feature this year. Four real-life leagues are supported in the new interactive league online mode. In case you're not familiar with the FIFA Lounge mode, the idea is that a group of you can play each other across multiple gaming sessions and have the game keep track of your results in a league table. Furthermore, you'll collect power-ups (or opponent power-downs) known as 'cheap shots' as you play that can be used to level the playing field in your favor before a subsequent match. In FIFA 06, these cards were allocated in such a way that losing players generally had more cheap shots available to them. In FIFA 07, however, the opposite is true, since winning players are almost always rewarded with better cheap shots. It's still possible to level the playing field to some extent before FIFA Lounge matches, but doing so is now achieved by awarding goals to a team before any cheap shots are activated. It's neat that when awarding goals to a team in this way, you can actually see a visual representation of how much it's likely to improve your chances of winning based on previous performances, but the system feels like a step in the wrong direction after last year's game. Winning a match against a superior player because you were able to bench his star forward or give all of his players a level of fatigue before kickoff is still rewarding, but beating that same player with a two-goal head start is a hollow victory at best. When you're not playing online or against your friends in the FIFA Lounge, you'll likely be putting your management skills to the test in FIFA 07's manager mode. The manager mode lets you assume control of any team in the game (or even one of your own creation) and then tasks you with leading them to glory while making decisions that can affect your club both on and off the field. After accepting a job at a team, your first duty will be to select a sponsor for the season. These sponsors won't replace the real ones on your uniforms when you play, but they're an important source of income, and you'll find that the sponsors offering you the most money are invariably the ones that will be the most difficult to please. In manager mode, picking and subsequently pleasing a sponsor is crucial for your club's finances. Next up will be an e-mail from your club's board of directors detailing their expectations for the season. Predictably, clubs that are currently enjoying a lot of success in real life expect it to continue, so choosing to manage a top-flight team can be more challenging than opting for one that's accustomed to midtable obscurity or relegation battles. The expectations are a little more varied in FIFA 07 than they were in FIFA 06, so in addition to achieving certain league and cup positions, you might find yourself tasked with improving club finances, reducing player salary bills, or extending the contracts of certain players. The board will also let you know which players are the fans' favorites, hoping that you'll find room for them in your starting 11 as a result. As in last year's game, your three main considerations in manager mode are keeping the fans happy, maintaining job security by keeping the board happy, and having good team chemistry. One of the new features for this year's manager mode is the player growth system, which lets you pluck upcoming players from your youth squad and then, by playing them alongside the first team, encouraging them to develop. Every player in your squad will gain experience points at the end of a match based on his performance, and you'll notice that the number of points awarded to young players is generally much higher than the number given to experienced pros. The flipside is that to nurture a star for the future, you have to spend multiple seasons fielding a player who isn't really good enough to be playing alongside the rest of your team. The other significant new feature in manager mode is the 'visual sim' option for matches. If for some reason you don't want to play a match yourself (we're not sure why that would ever be the case, frankly), the visual sim option lets you watch play-by-play commentary of the match and intervene with tactical decisions or by jumping in and assuming control at any time. The text-based commentary and match statistics that you have access to while in visual sim mode are adequate rather than impressive, and we can't help but wonder why there's no option to watch the game being played out using the regular, great-looking match engine. Fact is, the management portion of FIFA 07 works well in between matches, but it's not nearly deep enough to play purely as a management sim, which makes the option to 'quick sim' games and get a result instantly even more redundant than the aforementioned one. Quite why you'd ever want to play out your manager-mode matches in visual sim mode is a mystery. Regardless of the fact that the management in FIFA 07 isn't particularly deep (though it can be very engaging), to play the game without actually playing the matches yourself would be to miss out on some of the best soccer visuals that we've seen in a game to date. The players in FIFA 07 are instantly recognizable for the most part, but it's their animation that really stands out as a huge improvement over last year's game. In FIFA 06, the player animation was difficult to fault, but in FIFA 07, it's nigh on impossible--you'll see players controlling the ball with different parts of their bodies, you'll see them losing their balance and falling over occasionally, and you'll certainly notice them bumping into rather than clipping through each other when areas of the field get busy. FIFA 07, then, is a game that undoubtedly improves upon FIFA 06 in a number of ways, though it also has a few quirks of its own. This is an easy game to recommend if you have any interest in soccer, especially since you can keep up to date with the latest soccer news and results via a ticker tape along the bottom of the screen anytime you're online, but it's not a giant leap forward for the FIFA series in the same way that other recent iterations have been. FIFA 07 is a must-have if you missed out on FIFA 06, and it's definitely worth a look if you own last year's game and are ready for a change. This year, new intelligent A.I. Ensures that your 11 men on the pitch make realistic decisions, finding space and passing like professionals. A complete overhaul of the game engine now means that you have to employ real world tactics, make split-second decisions and think like a player in order to win matches. Watch as your players jostle and collide realistically while trying to win balls. Experience the realism of the world's superstars, who come to life with signature moves and authentic playing styles and a more sophisticated shooting mechanic that gives you greater control to produce finesse shots. Plus you now have the ability to apply topspin or backspin to the ball for more creative set-pieces. Atelier web security port scanner free download - Free Port Scanner, Avast Free Mac Security, Fast Port Scanner, and many more programs. Trusted Windows (PC) download Atelier Web Security Port Scanner 4.61. Virus-free and 100% clean download. Get Atelier Web Security Port Scanner alternative downloads. 
Atelier Web Security Port Scanner download page. Atelier Web Security Port Scanner by Jose Alberto Pupo Pascoa, download from Tools & Utilities subcategory. Best Buy There are thousands of pictures in many websites on Internet. And people often want to download thes. CounterPath's eyeBeam 1.5 a multimedia communicator designed to enhance the user's communi. With AV Voice Changer Software though, you can create and the nickvoices, either male or female. If you need a powerful yet simple-to-use download manager, don?t meddle, choose GetRight! FlashGet is specifically designed to address two of the biggest problems when downloading files: Spe. 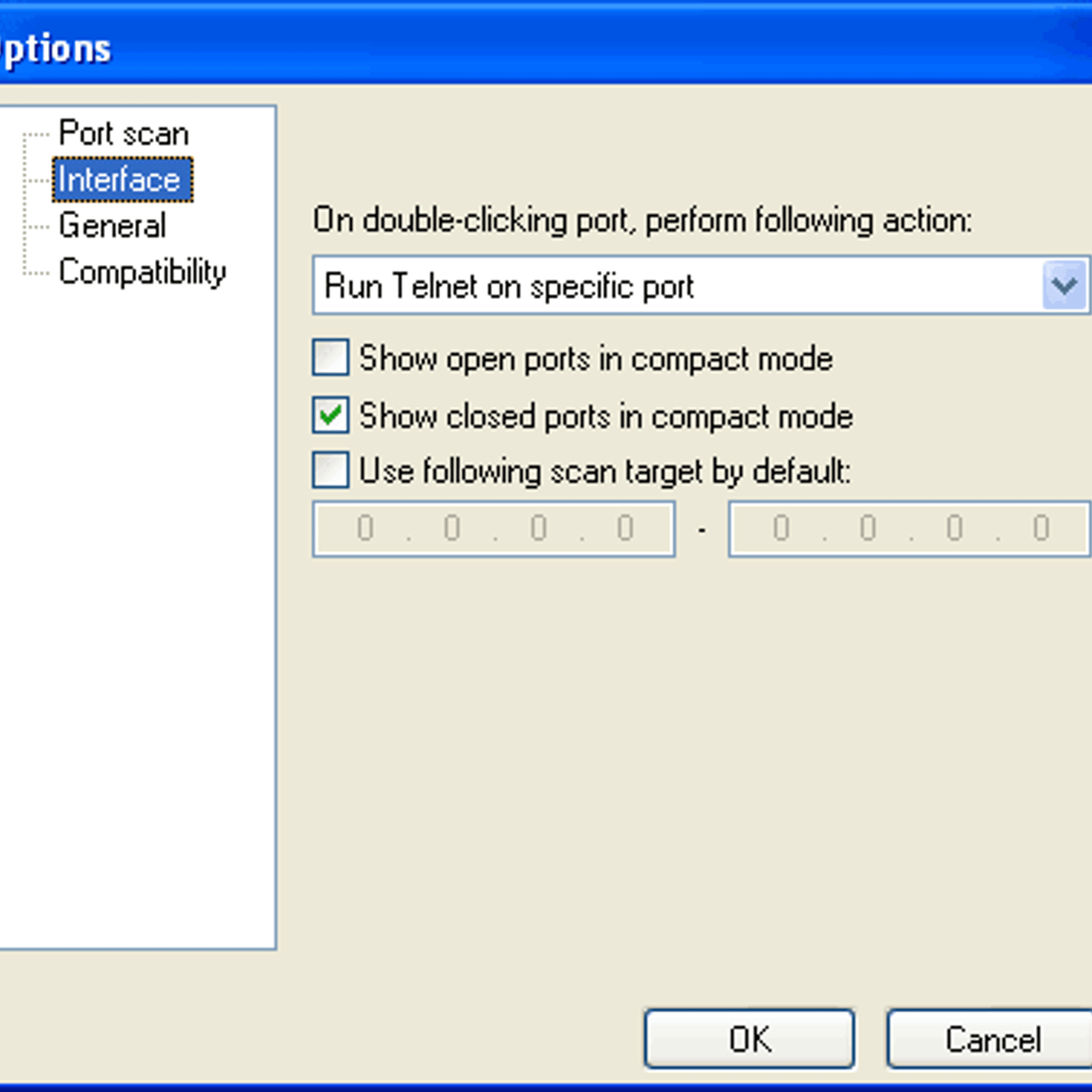 
|
AuthorWrite something about yourself. No need to be fancy, just an overview. Archives
February 2018
Categories |
 RSS Feed
RSS Feed
